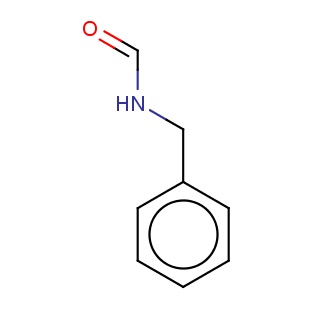
Binding information for 4o13_ligand_2_12.mol2(FDBF00182)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4o13_ligand_2_12.mol2 | 4o13 | 1 | -6.73 | N(C=O)Cc1ccccc1 | 10 |
Structure and binding mode of 4o13_ligand_2_12.mol2(FDBF00182)
Important binding residues for 4o13_ligand_2_12.mol2(FDBF00182)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4o13 | PHE193 | -1.21 | -0.56 | -1.77 | 0.64 | -1.13 |
| 4o13 | GLY217 | -0.06 | 0.03 | -0.03 | -0.29 | -0.32 |
| 4o13 | SER241 | -0.50 | 0.34 | -0.16 | -0.53 | -0.69 |
| 4o13 | VAL242 | -1.41 | -0.53 | -1.94 | 0.85 | -1.09 |
| 4o13 | ALA244 | -0.21 | -0.68 | -0.89 | 0.49 | -0.40 |
| 4o13 | SER275 | -0.83 | -1.15 | -1.98 | 1.47 | -0.51 |
| 4o13 | ILE309 | -0.45 | 0.22 | -0.23 | -0.25 | -0.48 |
| 4o13 | ILE351 | -0.68 | 0.01 | -0.67 | -0.11 | -0.78 |
| 4o13 | SER17 | -0.15 | 0.56 | 0.41 | -0.76 | -0.35 |
| 4o13 | TYR18 | -1.21 | -0.15 | -1.36 | -0.09 | -1.44 |