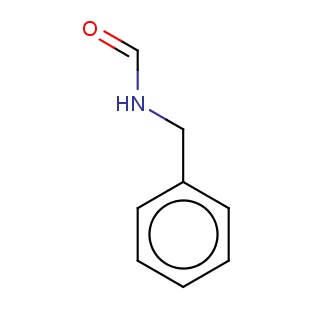
Binding information for 4r5x_ligand_2_1.mol2(FDBF00182)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4r5x_ligand_2_1.mol2 | 4r5x | 1 | -6.70 | N(C=O)Cc1ccccc1 | 10 |
Structure and binding mode of 4r5x_ligand_2_1.mol2(FDBF00182)
Important binding residues for 4r5x_ligand_2_1.mol2(FDBF00182)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4r5x | VAL459 | -1.37 | -1.63 | -3 | 0.50 | -2.50 |
| 4r5x | GLY460 | 0.34 | -3.68 | -3.34 | 1.25 | -2.08 |
| 4r5x | ALA461 | -0.81 | -0.88 | -1.69 | 0.88 | -0.81 |
| 4r5x | GLU519 | -0.28 | -1.73 | -2.01 | 0.41 | -1.60 |
| 4r5x | TYR575 | -1.46 | -0.30 | -1.76 | 0.79 | -0.97 |
| 4r5x | TYR580 | -1.44 | 0.91 | -0.53 | 0.01 | -0.53 |