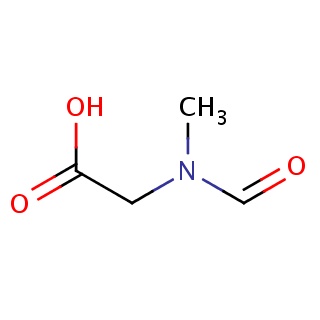
Binding information for 1nhv_ligand_2_0.mol2(FDBF00183)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1nhv_ligand_2_0.mol2 | 1nhv | 0.863636 | -5.13 | CN(CC(=O)O)C=O | 8 |
Structure and binding mode of 1nhv_ligand_2_0.mol2(FDBF00183)
Important binding residues for 1nhv_ligand_2_0.mol2(FDBF00183)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1nhv | HIS475 | -0.92 | -2.99 | -3.91 | 1.15 | -2.76 |
| 1nhv | SER476 | 0.07 | -10.72 | -10.65 | 7.29 | -3.35 |
| 1nhv | TYR477 | -0.67 | -1.96 | -2.63 | 0.11 | -2.51 |
| 1nhv | ARG501 | -0.20 | -20.36 | -20.56 | 20.16 | -0.40 |