Binding information for 2vl1_ligand.mol2(FDBF00183)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
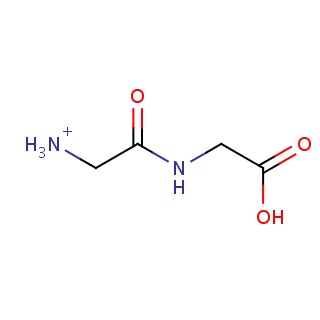
| 2vl1_ligand.mol2 | 2vl1 | 0.814815 | -6.81 | [NH3+]CC(=O)NCC(=O)O | 10 |
Structure and binding mode of 2vl1_ligand.mol2(FDBF00183)
Important binding residues for 2vl1_ligand.mol2(FDBF00183)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2vl1 | ASP116 | -0.04 | -6.70 | -6.74 | 6.05 | -0.69 |
| 2vl1 | ASP125 | 1.02 | -7.12 | -6.1 | -14.10 | -20.20 |
| 2vl1 | GLU159 | -0.90 | -19.80 | -20.7 | 8.87 | -11.83 |
| 2vl1 | GLU160 | 0.46 | -11.57 | -11.11 | -11.04 | -22.14 |
| 2vl1 | GLU224 | -0.03 | -4.38 | -4.41 | 3.53 | -0.88 |
| 2vl1 | TYR249 | -0.52 | 1.55 | 1.03 | -1.66 | -0.63 |
| 2vl1 | TRP251 | -0.42 | -0.03 | -0.45 | -0.09 | -0.53 |
| 2vl1 | ARG322 | 0.94 | -13.57 | -12.63 | 10.76 | -1.87 |
| 2vl1 | ALA395 | -1.11 | -1.61 | -2.72 | 1.82 | -0.90 |
| 2vl1 | GLY396 | 0.54 | -6.63 | -6.09 | 3.62 | -2.47 |
| 2vl1 | HIS397 | -0.88 | -0.95 | -1.83 | 0.10 | -1.74 |
| 2vl1 | ASP398 | -0.06 | -7.06 | -7.12 | 6.59 | -0.53 |
| 2vl1 | HIS421 | -0.67 | -1.02 | -1.69 | 0.95 | -0.74 |
| 2vl1 | GLU425 | -0.02 | -0.24 | -0.26 | -0.20 | -0.45 |
| 2vl1 | ASN309 | -0.66 | -2.27 | -2.93 | 2.45 | -0.48 |