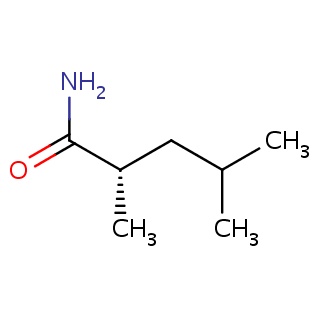
Binding information for 1y3g_ligand_4_680.mol2(FDBF00185)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1y3g_ligand_4_680.mol2 | 1y3g | 1 | -6.74 | C[C@H](C(=O)N)CC(C)C | 9 |
Structure and binding mode of 1y3g_ligand_4_680.mol2(FDBF00185)
Important binding residues for 1y3g_ligand_4_680.mol2(FDBF00185)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1y3g | VAL139 | -1.23 | 0.00 | -1.23 | -0.02 | -1.25 |
| 1y3g | GLU166 | -0.31 | 4.81 | 4.5 | -6.71 | -2.21 |
| 1y3g | LEU202 | -0.86 | -0.95 | -1.81 | 1.39 | -0.43 |
| 1y3g | ARG203 | -0.56 | -10.72 | -11.28 | 8.03 | -3.24 |