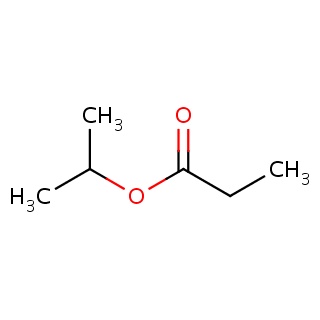
Binding information for 1p6d_ligand_4_1325.mol2(FDBF00192)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1p6d_ligand_4_1325.mol2 | 1p6d | 0.727273 | -6.51 | C(OC(=O)CC)(C)C | 8 |
Structure and binding mode of 1p6d_ligand_4_1325.mol2(FDBF00192)
Important binding residues for 1p6d_ligand_4_1325.mol2(FDBF00192)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1p6d | ASN55 | -0.21 | -0.12 | -0.33 | -0.17 | -0.50 |
| 1p6d | THR65 | -0.50 | -0.32 | -0.82 | 0.44 | -0.38 |
| 1p6d | PHE66 | -1.81 | 0.08 | -1.73 | 0.01 | -1.72 |
| 1p6d | HIS69 | -0.30 | -0.08 | -0.38 | -0.38 | -0.76 |
| 1p6d | PHE70 | -1.08 | -0.35 | -1.43 | 0.18 | -1.26 |
| 1p6d | TYR79 | -0.74 | 0.02 | -0.72 | 0.36 | -0.36 |
| 1p6d | ASP122 | -0.08 | -0.56 | -0.64 | -1.03 | -1.66 |
| 1p6d | THR133 | -1.16 | -1.81 | -2.97 | 1.06 | -1.91 |
| 1p6d | ASN134 | -0.77 | -2.52 | -3.29 | 2.29 | -1.00 |
| 1p6d | LEU135 | -0.53 | -0.10 | -0.63 | 0.14 | -0.49 |
| 1p6d | GLU146 | -0.26 | -0.58 | -0.84 | -2.94 | -3.78 |