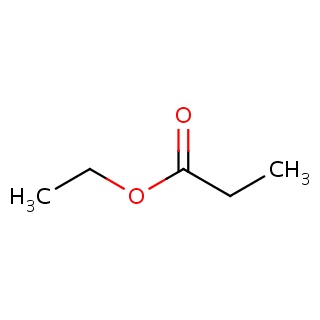
Binding information for 1p6e_ligand_3_208.mol2(FDBF00192)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1p6e_ligand_3_208.mol2 | 1p6e | 0.727273 | -5.97 | CCOC(=O)CC | 7 |
Structure and binding mode of 1p6e_ligand_3_208.mol2(FDBF00192)
Important binding residues for 1p6e_ligand_3_208.mol2(FDBF00192)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1p6e | THR65 | -0.44 | -0.15 | -0.59 | 0.24 | -0.35 |
| 1p6e | PHE66 | -1.61 | 0.01 | -1.6 | 0.10 | -1.50 |
| 1p6e | PHE70 | -0.70 | -0.18 | -0.88 | 0.15 | -0.73 |
| 1p6e | ASP122 | -0.03 | -0.09 | -0.12 | -0.44 | -0.56 |
| 1p6e | THR133 | -1.02 | -1.37 | -2.39 | 0.67 | -1.71 |
| 1p6e | ASN134 | -0.87 | -2.49 | -3.36 | 2.16 | -1.20 |
| 1p6e | LEU135 | -0.58 | -0.03 | -0.61 | 0.07 | -0.55 |
| 1p6e | HIS142 | -0.72 | 0.79 | 0.07 | -0.61 | -0.54 |
| 1p6e | GLU146 | -0.12 | -0.01 | -0.13 | -1.47 | -1.59 |