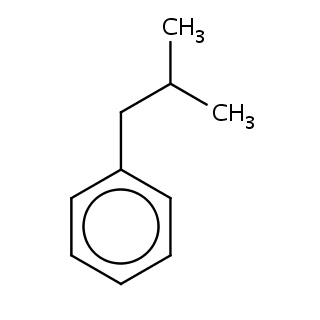
Binding information for 184l_ligand.mol2(FDBF00193)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 184l_ligand.mol2 | 184l | 1 | -7.92 | c1(ccccc1)CC(C)C | 11 |
Structure and binding mode of 184l_ligand.mol2(FDBF00193)
Important binding residues for 184l_ligand.mol2(FDBF00193)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 184l | ILE78 | -0.74 | 0.04 | -0.7 | -0.03 | -0.73 |
| 184l | LEU84 | -1.60 | -0.24 | -1.84 | 0.17 | -1.66 |
| 184l | VAL87 | -0.80 | 0.20 | -0.6 | -0.25 | -0.85 |
| 184l | TYR88 | -0.65 | -0.16 | -0.81 | 0.24 | -0.57 |
| 184l | ALA99 | -1.21 | -0.59 | -1.8 | 0.35 | -1.45 |
| 184l | MET102 | -1.12 | -0.03 | -1.15 | 0.15 | -1.00 |
| 184l | VAL103 | -0.50 | -0.00 | -0.5 | -0.07 | -0.57 |
| 184l | VAL111 | -0.75 | 0.06 | -0.69 | 0.06 | -0.63 |
| 184l | PHE114 | -0.87 | -0.01 | -0.88 | 0.19 | -0.69 |
| 184l | SER117 | -0.44 | -0.01 | -0.45 | 0.12 | -0.33 |
| 184l | LEU118 | -0.60 | -0.08 | -0.68 | 0.11 | -0.56 |
| 184l | LEU121 | -0.84 | 0.04 | -0.8 | -0.02 | -0.82 |
| 184l | PHE153 | -0.69 | -0.13 | -0.82 | 0.42 | -0.40 |