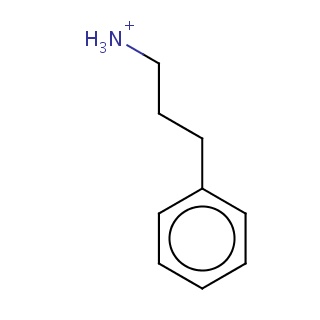
Binding information for 4jbs_ligand_3_0.mol2(FDBF00194)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4jbs_ligand_3_0.mol2 | 4jbs | 1 | -6.99 | C([NH3+])CCc1ccccc1 | 10 |
Structure and binding mode of 4jbs_ligand_3_0.mol2(FDBF00194)
Important binding residues for 4jbs_ligand_3_0.mol2(FDBF00194)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4jbs | GLU200 | 0.58 | -54.31 | -53.73 | 45.28 | -8.45 |
| 4jbs | PRO201 | -0.42 | 0.16 | -0.26 | -0.07 | -0.33 |
| 4jbs | PRO333 | -0.53 | 1.90 | 1.37 | -1.88 | -0.52 |
| 4jbs | ALA335 | -0.53 | -3.07 | -3.6 | 3.14 | -0.46 |
| 4jbs | MET336 | -1.10 | -0.84 | -1.94 | 0.46 | -1.49 |
| 4jbs | GLU337 | 2.39 | -53.16 | -50.77 | 39.98 | -10.79 |
| 4jbs | GLU371 | -0.17 | -29.03 | -29.2 | 28.73 | -0.47 |
| 4jbs | ILE389 | -0.08 | -16.96 | -17.04 | 16.67 | -0.37 |
| 4jbs | GLU393 | -0.58 | -55.62 | -56.2 | 48.15 | -8.05 |
| 4jbs | PHE450 | -1.76 | 0.83 | -0.93 | -0.33 | -1.25 |
| 4jbs | TYR455 | -0.56 | -0.95 | -1.51 | 0.92 | -0.59 |