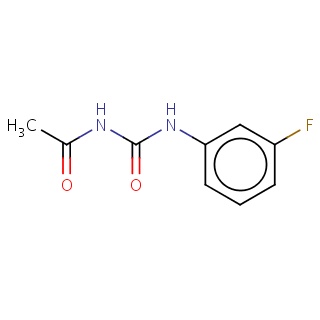
Binding information for 3ctj_ligand_2_4.mol2(FDBF05035)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3ctj_ligand_2_4.mol2 | 3ctj | 0.556818 | -7.20 | CC(=O)NC(=O)Nc1cc(F)ccc1 | 14 |
Structure and binding mode of 3ctj_ligand_2_4.mol2(FDBF05035)
Important binding residues for 3ctj_ligand_2_4.mol2(FDBF05035)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3ctj | VAL1092 | -0.16 | -0.26 | -0.42 | 0.11 | -0.31 |
| 3ctj | LYS1110 | -0.67 | -2.85 | -3.52 | 2.86 | -0.67 |
| 3ctj | MET1131 | -1.26 | -2.04 | -3.3 | 1.81 | -1.48 |
| 3ctj | LEU1140 | -1.23 | -0.47 | -1.7 | 0.43 | -1.28 |
| 3ctj | LEU1157 | -1.21 | -0.22 | -1.43 | 0.06 | -1.37 |
| 3ctj | MET1211 | -0.56 | -0.36 | -0.92 | 0.31 | -0.62 |
| 3ctj | ALA1221 | -0.89 | -0.86 | -1.75 | 0.35 | -1.39 |
| 3ctj | ASP1222 | -1.23 | -3.81 | -5.04 | 3.84 | -1.20 |
| 3ctj | PHE1223 | -2.18 | -0.41 | -2.59 | 2.23 | -0.35 |