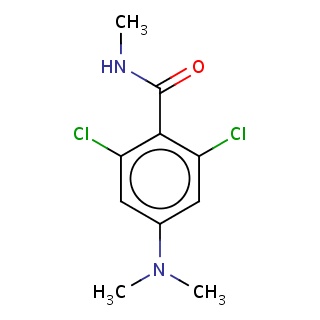
Binding information for 4oiv_ligand_3_0.mol2(FDBF05559)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4oiv_ligand_3_0.mol2 | 4oiv | 0.607143 | -7.48 | C(=O)(NC)c1c(Cl)cc(cc1Cl)N(C)C | 15 |
Structure and binding mode of 4oiv_ligand_3_0.mol2(FDBF05559)
Important binding residues for 4oiv_ligand_3_0.mol2(FDBF05559)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4oiv | THR292 | -1.30 | 0.17 | -1.13 | 0.31 | -0.82 |
| 4oiv | ALA295 | -1.46 | -0.20 | -1.66 | -0.26 | -1.91 |
| 4oiv | THR296 | -0.69 | -0.10 | -0.79 | 0.26 | -0.53 |
| 4oiv | VAL299 | -0.44 | 0.29 | -0.15 | -0.38 | -0.53 |
| 4oiv | VAL329 | -0.38 | -0.10 | -0.48 | 0.09 | -0.39 |
| 4oiv | MET332 | -1.56 | -0.45 | -2.01 | 0.47 | -1.54 |
| 4oiv | PHE333 | -1.04 | -0.58 | -1.62 | 0.72 | -0.90 |
| 4oiv | MET369 | -0.49 | 0.20 | -0.29 | -0.16 | -0.44 |
| 4oiv | HIS451 | -0.66 | -1.25 | -1.91 | 1.15 | -0.76 |
| 4oiv | ALA452 | -1.25 | -0.66 | -1.91 | 0.67 | -1.24 |
| 4oiv | MET454 | -0.14 | -0.43 | -0.57 | 0.19 | -0.37 |
| 4oiv | LEU455 | -2.83 | -1.11 | -3.94 | 1.90 | -2.04 |
| 4oiv | MET456 | -1.66 | -1.23 | -2.89 | 1.44 | -1.46 |
| 4oiv | TRP458 | -0.89 | -1.22 | -2.11 | 1.70 | -0.41 |
| 4oiv | ARG459 | -0.46 | 3.22 | 2.76 | -3.18 | -0.41 |