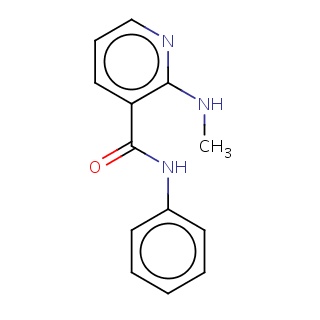
Binding information for 2p2i_ligand_4_16.mol2(FDBF06004)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2p2i_ligand_4_16.mol2 | 2p2i | 0.697917 | -8.49 | c1cc(ccc1)NC(=O)c1cccnc1NC | 17 |
Structure and binding mode of 2p2i_ligand_4_16.mol2(FDBF06004)
Important binding residues for 2p2i_ligand_4_16.mol2(FDBF06004)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2p2i | VAL848 | -1.19 | -0.05 | -1.24 | -0.02 | -1.26 |
| 2p2i | ALA866 | -0.98 | -0.66 | -1.64 | 0.60 | -1.04 |
| 2p2i | LYS868 | -2.23 | 2.10 | -0.13 | -0.96 | -1.08 |
| 2p2i | GLU885 | -0.80 | -8.64 | -9.44 | 8.60 | -0.84 |
| 2p2i | LEU889 | -1.15 | -0.35 | -1.5 | 0.35 | -1.15 |
| 2p2i | VAL898 | -0.33 | -0.07 | -0.4 | 0.06 | -0.33 |
| 2p2i | VAL899 | -1.14 | -0.38 | -1.52 | 0.32 | -1.20 |
| 2p2i | VAL914 | -0.74 | -0.48 | -1.22 | 0.18 | -1.05 |
| 2p2i | ILE915 | -0.27 | -0.12 | -0.39 | 0.07 | -0.32 |
| 2p2i | VAL916 | -1.77 | -0.12 | -1.89 | -0.06 | -1.96 |
| 2p2i | ILE1044 | -0.46 | -0.11 | -0.57 | -0.10 | -0.67 |
| 2p2i | CYS1045 | -1.30 | -3.29 | -4.59 | 1.35 | -3.23 |
| 2p2i | ASP1046 | -1.97 | -0.64 | -2.61 | 0.44 | -2.16 |
| 2p2i | PHE1047 | -1.38 | -0.27 | -1.65 | 0.52 | -1.13 |