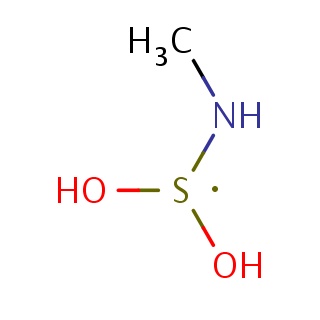
Binding information for 4jxs_ligand_1_1.mol2(FDBF06441)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4jxs_ligand_1_1.mol2 | 4jxs | 0.416667 | -5.50 | CN[S](O)O | 5 |
Structure and binding mode of 4jxs_ligand_1_1.mol2(FDBF06441)
Important binding residues for 4jxs_ligand_1_1.mol2(FDBF06441)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4jxs | SER64 | 1.05 | -7.42 | -6.37 | -1.69 | -8.05 |
| 4jxs | GLN120 | -0.40 | -0.30 | -0.7 | 0.20 | -0.50 |
| 4jxs | ASN152 | 0.47 | -9.60 | -9.13 | 5.33 | -3.81 |
| 4jxs | TYR221 | -0.91 | -0.09 | -1 | 0.33 | -0.67 |
| 4jxs | ALA318 | -0.04 | -4.25 | -4.29 | 3.53 | -0.77 |
| 4jxs | THR319 | -0.49 | -0.01 | -0.5 | 0.17 | -0.33 |