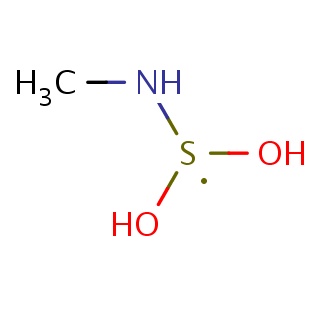
Binding information for 2jsd_ligand_1_3.mol2(FDBF06441)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2jsd_ligand_1_3.mol2 | 2jsd | 0.416667 | -5.47 | [S](O)(O)NC | 5 |
Structure and binding mode of 2jsd_ligand_1_3.mol2(FDBF06441)
Important binding residues for 2jsd_ligand_1_3.mol2(FDBF06441)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2jsd | ASP183 | -0.05 | -34.39 | -34.44 | 33.94 | -0.49 |
| 2jsd | GLY187 | -0.41 | -2.75 | -3.16 | 2.81 | -0.36 |
| 2jsd | THR188 | -0.77 | 1.09 | 0.32 | -1.06 | -0.74 |
| 2jsd | ALA190 | 0.07 | -8.79 | -8.72 | 6.15 | -2.56 |
| 2jsd | HIS191 | -0.41 | -0.45 | -0.86 | 0.25 | -0.61 |
| 2jsd | ASP206 | -0.02 | -30.01 | -30.03 | 29.63 | -0.40 |
| 2jsd | GLU209 | -0.02 | -31.17 | -31.19 | 30.74 | -0.46 |
| 2jsd | GLU227 | -0.23 | -27.97 | -28.2 | 27.80 | -0.40 |
| 2jsd | HIS236 | -0.47 | -11.41 | -11.88 | 11.23 | -0.64 |