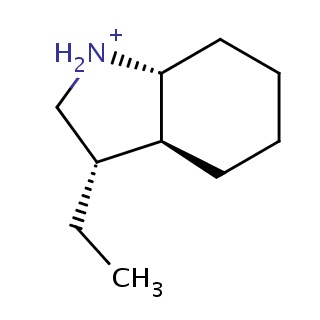
Binding information for 1wbs_ligand_2_12.mol2(FDBF00010)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1wbs_ligand_2_12.mol2 | 1wbs | 1 | -7.64 | C([C@@H]1C[NH2+][C@@H]2CCCC[C@@H]12)C | 11 |
Structure and binding mode of 1wbs_ligand_2_12.mol2(FDBF00010)
Important binding residues for 1wbs_ligand_2_12.mol2(FDBF00010)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1wbs | VAL38 | -0.86 | 0.18 | -0.68 | -0.21 | -0.88 |
| 1wbs | ALA51 | -0.89 | -1.13 | -2.02 | 0.67 | -1.34 |
| 1wbs | VAL52 | -0.55 | 0.03 | -0.52 | -0.02 | -0.53 |
| 1wbs | LYS53 | -2.66 | -3.93 | -6.59 | 3.35 | -3.25 |
| 1wbs | ILE84 | -0.85 | -0.13 | -0.98 | 0.02 | -0.96 |
| 1wbs | LEU104 | -1.40 | -0.93 | -2.33 | 0.52 | -1.82 |
| 1wbs | VAL105 | -0.50 | 0.37 | -0.13 | -0.30 | -0.43 |
| 1wbs | THR106 | -1.18 | -0.12 | -1.3 | 0.04 | -1.27 |
| 1wbs | LEU167 | -0.52 | -0.14 | -0.66 | -0.03 | -0.69 |
| 1wbs | PHE169 | -1.37 | -0.07 | -1.44 | 0.52 | -0.92 |