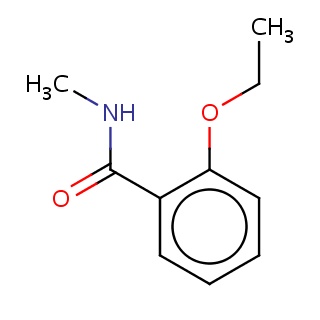
Binding information for 2v12_ligand_5_4200.mol2(FDBF07586)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2v12_ligand_5_4200.mol2 | 2v12 | 0.830508 | -6.91 | C(C)Oc1ccccc1C(=O)NC | 13 |
Structure and binding mode of 2v12_ligand_5_4200.mol2(FDBF07586)
Important binding residues for 2v12_ligand_5_4200.mol2(FDBF07586)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2v12 | GLN19 | -1.35 | 0.37 | -0.98 | 0.30 | -0.68 |
| 2v12 | VAL36 | -0.55 | 0.26 | -0.29 | -0.25 | -0.54 |
| 2v12 | PRO118 | -0.79 | -0.89 | -1.68 | 1.06 | -0.61 |
| 2v12 | PHE119 | -0.62 | -0.11 | -0.73 | 0.21 | -0.52 |
| 2v12 | LEU121 | -0.50 | -0.17 | -0.67 | 0.32 | -0.35 |
| 2v12 | ALA122 | -0.58 | 0.16 | -0.42 | -0.15 | -0.57 |
| 2v12 | PHE124 | -1.07 | -0.28 | -1.35 | 0.41 | -0.95 |
| 2v12 | ALA229 | -0.52 | -0.34 | -0.86 | 0.46 | -0.40 |