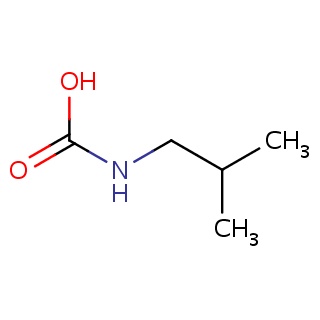
Binding information for 9hvp_ligand_2_144.mol2(FDBF07594)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 9hvp_ligand_2_144.mol2 | 9hvp | 1 | -6.01 | C(NC(=O)O)C(C)C | 8 |
Structure and binding mode of 9hvp_ligand_2_144.mol2(FDBF07594)
Important binding residues for 9hvp_ligand_2_144.mol2(FDBF07594)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 9hvp | ARG8 | -0.15 | -14.21 | -14.36 | 13.98 | -0.38 |
| 9hvp | ILE50 | -0.74 | -1.23 | -1.97 | 1.18 | -0.79 |
| 9hvp | GLY27 | -0.39 | 2.71 | 2.32 | -2.66 | -0.34 |
| 9hvp | ALA28 | -0.29 | -6.27 | -6.56 | 2.82 | -3.75 |
| 9hvp | ASP29 | -0.30 | 26.50 | 26.2 | -27.90 | -1.70 |
| 9hvp | LYS45 | -0.03 | -32.13 | -32.16 | 31.85 | -0.31 |
| 9hvp | ILE47 | -0.26 | -2.14 | -2.4 | 1.94 | -0.46 |
| 9hvp | ILE84 | -0.21 | -0.35 | -0.56 | 0.22 | -0.34 |