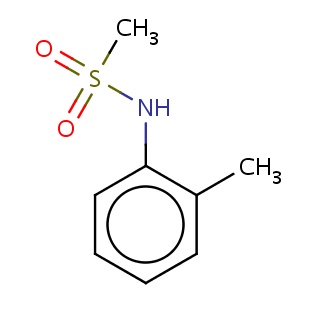
Binding information for 3exo_ligand_1_1.mol2(FDBF07600)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3exo_ligand_1_1.mol2 | 3exo | 0.583333 | -6.94 | N(S(=O)(=O)C)c1c(cccc1)C | 12 |
Structure and binding mode of 3exo_ligand_1_1.mol2(FDBF07600)
Important binding residues for 3exo_ligand_1_1.mol2(FDBF07600)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3exo | GLN12 | -1.07 | 0.47 | -0.6 | -0.29 | -0.89 |
| 3exo | GLY13 | -0.80 | -0.36 | -1.16 | 0.43 | -0.73 |
| 3exo | LEU30 | -1.23 | -2.30 | -3.53 | 2.32 | -1.21 |
| 3exo | TYR71 | -0.84 | -0.04 | -0.88 | 0.41 | -0.47 |
| 3exo | ILE110 | -0.73 | 0.01 | -0.72 | -0.05 | -0.78 |
| 3exo | TRP115 | -1.34 | 0.48 | -0.86 | -0.18 | -1.04 |
| 3exo | ILE118 | -0.58 | -0.03 | -0.61 | -0.00 | -0.62 |
| 3exo | GLY230 | -0.56 | -5.00 | -5.56 | 3.30 | -2.26 |