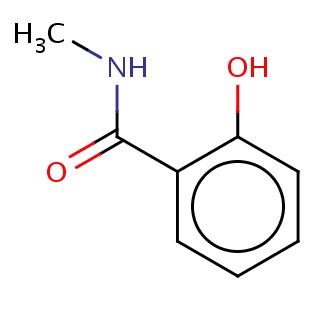
Binding information for 1r6g_ligand_2_15.mol2(FDBF07614)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1r6g_ligand_2_15.mol2 | 1r6g | 0.773585 | -6.82 | c1cc(c(cc1)O)C(=O)NC | 11 |
Structure and binding mode of 1r6g_ligand_2_15.mol2(FDBF07614)
Important binding residues for 1r6g_ligand_2_15.mol2(FDBF07614)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1r6g | PHE272 | -1.44 | -0.45 | -1.89 | 0.50 | -1.39 |
| 1r6g | THR273 | -0.50 | -0.12 | -0.62 | 0.20 | -0.43 |
| 1r6g | ILE276 | -1.01 | -0.07 | -1.08 | -0.06 | -1.14 |
| 1r6g | MET310 | -0.69 | -0.39 | -1.08 | 0.40 | -0.68 |
| 1r6g | MET313 | -0.44 | 0.19 | -0.25 | -0.14 | -0.38 |
| 1r6g | LEU330 | -0.39 | 0.11 | -0.28 | -0.13 | -0.41 |
| 1r6g | LEU341 | -0.67 | 0.81 | 0.14 | -0.70 | -0.56 |
| 1r6g | LEU346 | -1.77 | -0.58 | -2.35 | 0.24 | -2.12 |
| 1r6g | HIS435 | -0.81 | -0.96 | -1.77 | 0.93 | -0.84 |