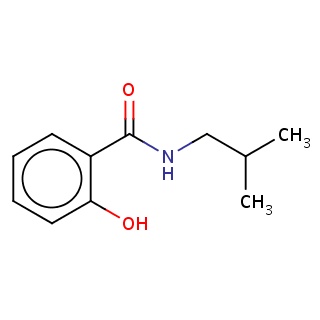
Binding information for 2v13_ligand_5_4901.mol2(FDBF07614)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2v13_ligand_5_4901.mol2 | 2v13 | 0.762712 | -7.40 | Oc1ccccc1C(=O)NCC(C)C | 14 |
Structure and binding mode of 2v13_ligand_5_4901.mol2(FDBF07614)
Important binding residues for 2v13_ligand_5_4901.mol2(FDBF07614)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2v13 | VAL36 | -0.43 | 0.06 | -0.37 | -0.18 | -0.55 |
| 2v13 | TYR83 | -1.16 | 0.08 | -1.08 | 0.31 | -0.77 |
| 2v13 | PRO118 | -0.66 | -0.85 | -1.51 | 0.87 | -0.64 |
| 2v13 | PHE119 | -1.03 | -0.22 | -1.25 | 0.39 | -0.86 |
| 2v13 | LEU121 | -0.60 | -0.15 | -0.75 | 0.25 | -0.50 |
| 2v13 | ALA122 | -0.50 | -0.04 | -0.54 | -0.10 | -0.63 |
| 2v13 | PHE124 | -1.42 | -0.55 | -1.97 | 1.01 | -0.96 |
| 2v13 | VAL127 | -0.59 | 0.07 | -0.52 | -0.24 | -0.76 |