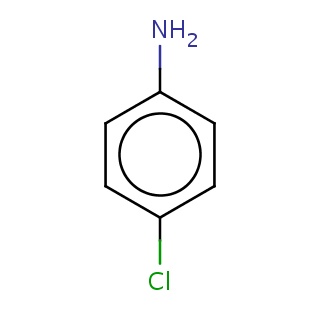
Binding information for 3idp_ligand_1_2.mol2(FDBF07625)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3idp_ligand_1_2.mol2 | 3idp | 0.512821 | -6.54 | Nc1ccc(Cl)cc1 | 8 |
Structure and binding mode of 3idp_ligand_1_2.mol2(FDBF07625)
Important binding residues for 3idp_ligand_1_2.mol2(FDBF07625)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3idp | GLU501 | -0.26 | -7.38 | -7.64 | 6.92 | -0.73 |
| 3idp | VAL504 | -0.63 | -0.40 | -1.03 | 0.45 | -0.58 |
| 3idp | LEU505 | -1.16 | 0.59 | -0.57 | -0.49 | -1.07 |
| 3idp | ILE513 | -0.37 | 0.03 | -0.34 | -0.02 | -0.36 |
| 3idp | LEU567 | -0.48 | 0.86 | 0.38 | -0.82 | -0.44 |
| 3idp | ILE592 | -0.31 | -0.39 | -0.7 | 0.35 | -0.35 |
| 3idp | GLY593 | -0.68 | -0.68 | -1.36 | 0.34 | -1.02 |
| 3idp | ASP594 | -1.28 | 0.33 | -0.95 | 0.12 | -0.84 |