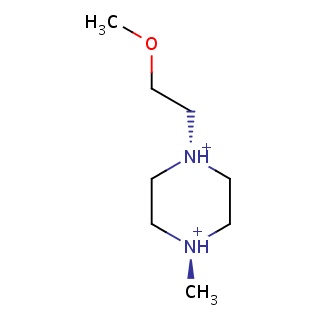
Binding information for 5e1s_ligand_4_55.mol2(FDBF07628)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 5e1s_ligand_4_55.mol2 | 5e1s | 0.488889 | -5.02 | C(OC)C[N@@H+]1CC[N@@H+](C)CC1 | 11 |
Structure and binding mode of 5e1s_ligand_4_55.mol2(FDBF07628)
Important binding residues for 5e1s_ligand_4_55.mol2(FDBF07628)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 5e1s | GLU1012 | -0.04 | -36.57 | -36.61 | 36.31 | -0.31 |
| 5e1s | GLY1082 | -0.54 | 0.79 | 0.25 | -1.02 | -0.77 |
| 5e1s | ASP1083 | -0.10 | -39.48 | -39.58 | 39.20 | -0.38 |
| 5e1s | PRO1099 | -0.56 | 1.04 | 0.48 | -1.06 | -0.58 |
| 5e1s | VAL1140 | -0.05 | -34.07 | -34.12 | 33.74 | -0.39 |