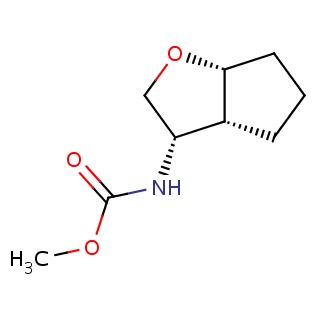
Binding information for 4dfg_ligand_1_10.mol2(FDBF00706)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4dfg_ligand_1_10.mol2 | 4dfg | 0.604938 | -6.49 | N(C(=O)OC)[C@@H]1CO[C@@H]2CCC[C@H]12 | 13 |
Structure and binding mode of 4dfg_ligand_1_10.mol2(FDBF00706)
Important binding residues for 4dfg_ligand_1_10.mol2(FDBF00706)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4dfg | ARG8 | -0.91 | -3.27 | -4.18 | 2.85 | -1.33 |
| 4dfg | ILE50 | -0.47 | 0.03 | -0.44 | -0.04 | -0.48 |
| 4dfg | VAL82 | -0.44 | -0.14 | -0.58 | 0.03 | -0.55 |
| 4dfg | ALA28 | -0.96 | -0.87 | -1.83 | 0.15 | -1.69 |
| 4dfg | ILE47 | -1.06 | 0.51 | -0.55 | -0.32 | -0.87 |
| 4dfg | GLY49 | -0.62 | -0.20 | -0.82 | 0.33 | -0.50 |