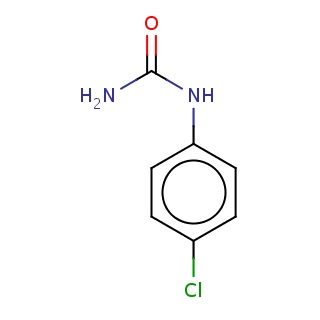
Binding information for 1uwh_ligand_1_0.mol2(FDBF00873)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1uwh_ligand_1_0.mol2 | 1uwh | 0.902439 | -6.90 | C(=O)(Nc1ccc(cc1)Cl)N | 11 |
Structure and binding mode of 1uwh_ligand_1_0.mol2(FDBF00873)
Important binding residues for 1uwh_ligand_1_0.mol2(FDBF00873)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1uwh | GLU500 | -0.93 | -12.58 | -13.51 | 11.01 | -2.50 |
| 1uwh | VAL503 | -0.66 | 0.09 | -0.57 | -0.04 | -0.61 |
| 1uwh | LEU504 | -1.49 | 0.21 | -1.28 | -0.26 | -1.55 |
| 1uwh | LEU513 | -0.53 | -0.32 | -0.85 | 0.23 | -0.62 |
| 1uwh | HIS573 | -0.80 | -0.33 | -1.13 | 0.61 | -0.52 |
| 1uwh | GLY592 | -0.90 | -1.77 | -2.67 | 1.08 | -1.60 |
| 1uwh | ASP593 | -2.29 | -1.98 | -4.27 | 2.04 | -2.23 |