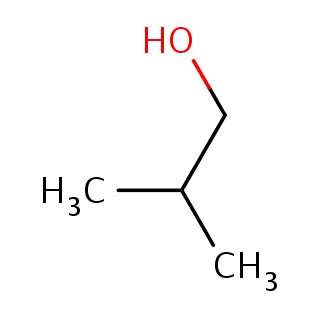
Binding information for 3guz_ligand_1_1.mol2(FDBF00018)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3guz_ligand_1_1.mol2 | 3guz | 1 | -6.03 | C(O)C(C)C | 5 |
Structure and binding mode of 3guz_ligand_1_1.mol2(FDBF00018)
Important binding residues for 3guz_ligand_1_1.mol2(FDBF00018)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3guz | PRO28 | -0.94 | -0.15 | -1.09 | 0.61 | -0.47 |
| 3guz | THR29 | -0.72 | 0.36 | -0.36 | -0.04 | -0.40 |
| 3guz | MET30 | -0.98 | -0.05 | -1.03 | 0.17 | -0.86 |
| 3guz | ASN58 | -0.38 | -1.02 | -1.4 | 0.28 | -1.12 |
| 3guz | VAL130 | -0.62 | 0.16 | -0.46 | -0.10 | -0.56 |
| 3guz | ILE133 | -0.47 | -0.33 | -0.8 | 0.13 | -0.67 |
| 3guz | VAL134 | -0.65 | 0.04 | -0.61 | 0.06 | -0.54 |