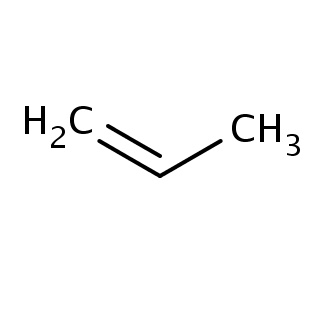
Binding information for 3dzt_ligand_1_9.mol2(FDBF00022)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 3dzt_ligand_1_9.mol2 | 3dzt | 1 | -5.91 | C=CC | 3 |
Structure and binding mode of 3dzt_ligand_1_9.mol2(FDBF00022)
Important binding residues for 3dzt_ligand_1_9.mol2(FDBF00022)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 3dzt | TRP37 | -1.49 | 0.30 | -1.19 | 0.27 | -0.92 |
| 3dzt | PHE50 | -0.54 | -0.11 | -0.65 | 0.26 | -0.40 |
| 3dzt | VAL54 | -0.36 | 0.00 | -0.36 | -0.08 | -0.44 |