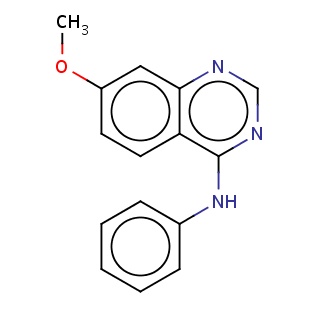
Binding information for 1di9_ligand_3_1.mol2(FDBF01207)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1di9_ligand_3_1.mol2 | 1di9 | 0.744444 | -7.69 | O(C)c1ccc2c(ncnc2Nc2ccccc2)c1 | 19 |
Structure and binding mode of 1di9_ligand_3_1.mol2(FDBF01207)
Important binding residues for 1di9_ligand_3_1.mol2(FDBF01207)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1di9 | VAL30 | -1.54 | -0.14 | -1.68 | 0.16 | -1.52 |
| 1di9 | VAL38 | -1.46 | -0.17 | -1.63 | 0.05 | -1.59 |
| 1di9 | ALA51 | -1.05 | -0.24 | -1.29 | 0.21 | -1.07 |
| 1di9 | LYS53 | -1.32 | -2.04 | -3.36 | 2.34 | -1.02 |
| 1di9 | LEU104 | -0.32 | 0.07 | -0.25 | -0.08 | -0.32 |
| 1di9 | LEU108 | -1.37 | -1.18 | -2.55 | 0.60 | -1.95 |
| 1di9 | MET109 | -1.25 | -3.21 | -4.46 | 2.08 | -2.37 |
| 1di9 | LEU167 | -1.07 | -0.79 | -1.86 | 1.22 | -0.65 |