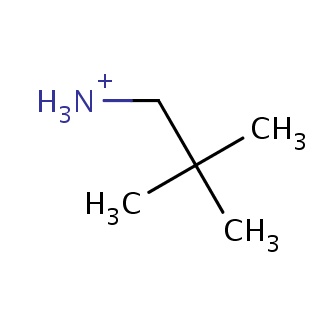
Binding information for 2ajb_ligand_1_0.mol2(FDBF00025)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2ajb_ligand_1_0.mol2 | 2ajb | 1 | -6.25 | C(C)(C)(C)C[NH3+] | 6 |
Structure and binding mode of 2ajb_ligand_1_0.mol2(FDBF00025)
Important binding residues for 2ajb_ligand_1_0.mol2(FDBF00025)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2ajb | VAL202 | -0.02 | -19.43 | -19.45 | 19.04 | -0.41 |
| 2ajb | GLU205 | 1.23 | -49.09 | -47.86 | 43.37 | -4.49 |
| 2ajb | GLU206 | 1.81 | -63.04 | -61.23 | 54.21 | -7.02 |
| 2ajb | TYR662 | -0.52 | 14.00 | 13.48 | -14.62 | -1.14 |
| 2ajb | TYR666 | -1.26 | 0.72 | -0.54 | -0.61 | -1.16 |
| 2ajb | ASP708 | -0.01 | -29.87 | -29.88 | 29.52 | -0.36 |