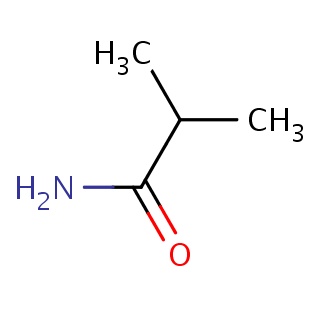
Binding information for 2j62_ligand_1_0.mol2(FDBF00026)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2j62_ligand_1_0.mol2 | 2j62 | 1 | -6.47 | CC(C)C(=O)N | 6 |
Structure and binding mode of 2j62_ligand_1_0.mol2(FDBF00026)
Important binding residues for 2j62_ligand_1_0.mol2(FDBF00026)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2j62 | ASP297 | -0.63 | -7.35 | -7.98 | 7.27 | -0.71 |
| 2j62 | VAL331 | -0.48 | 0.04 | -0.44 | 0.02 | -0.43 |
| 2j62 | TYR335 | -1.49 | 0.07 | -1.42 | 0.24 | -1.18 |
| 2j62 | VAL370 | -0.32 | 0.35 | 0.03 | -0.39 | -0.36 |
| 2j62 | TRP394 | -1.81 | -1.25 | -3.06 | 1.13 | -1.93 |
| 2j62 | ASN396 | -0.44 | -2.81 | -3.25 | 0.92 | -2.33 |