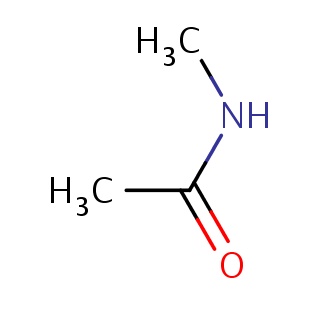
Binding information for 2g1r_ligand_1_4.mol2(FDBF00027)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2g1r_ligand_1_4.mol2 | 2g1r | 1 | -5.95 | CNC(=O)C | 5 |
Structure and binding mode of 2g1r_ligand_1_4.mol2(FDBF00027)
Important binding residues for 2g1r_ligand_1_4.mol2(FDBF00027)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2g1r | THR13 | -0.71 | 0.04 | -0.67 | 0.28 | -0.39 |
| 2g1r | GLN14 | 0.50 | -2.53 | -2.03 | 0.76 | -1.26 |
| 2g1r | TYR15 | 0.07 | -2.06 | -1.99 | 0.60 | -1.39 |
| 2g1r | VAL31 | -1.10 | -0.21 | -1.31 | 0.06 | -1.25 |
| 2g1r | GLY223 | -0.48 | -1.89 | -2.37 | 1.87 | -0.51 |
| 2g1r | ALA224 | -0.57 | -0.21 | -0.78 | 0.36 | -0.42 |
| 2g1r | SER225 | -0.65 | -0.40 | -1.05 | 0.62 | -0.42 |