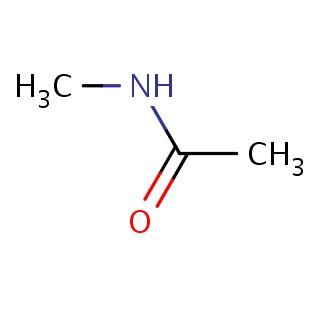
Binding information for 5abe_ligand_1_0.mol2(FDBF00027)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 5abe_ligand_1_0.mol2 | 5abe | 1 | -5.95 | CC(=O)NC | 5 |
Structure and binding mode of 5abe_ligand_1_0.mol2(FDBF00027)
Important binding residues for 5abe_ligand_1_0.mol2(FDBF00027)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 5abe | ASP242 | -0.40 | -8.31 | -8.71 | 8.37 | -0.34 |
| 5abe | TYR282 | -1.88 | 0.65 | -1.23 | 0.06 | -1.18 |
| 5abe | THR310 | -0.56 | -0.06 | -0.62 | 0.07 | -0.55 |
| 5abe | VAL314 | -0.29 | 0.55 | 0.26 | -0.57 | -0.31 |
| 5abe | TRP337 | -1.82 | -1.02 | -2.84 | 1.42 | -1.41 |
| 5abe | ASN339 | -0.30 | -3.93 | -4.23 | 1.53 | -2.69 |