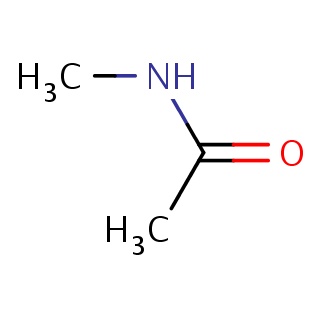
Binding information for 1a7c_ligand_1_0.mol2(FDBF00027)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1a7c_ligand_1_0.mol2 | 1a7c | 1 | -5.91 | N(C(=O)C)C | 5 |
Structure and binding mode of 1a7c_ligand_1_0.mol2(FDBF00027)
Important binding residues for 1a7c_ligand_1_0.mol2(FDBF00027)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1a7c | GLY173 | -0.58 | -2.61 | -3.19 | 2.17 | -1.02 |
| 1a7c | GLN174 | -0.60 | -0.45 | -1.05 | 0.55 | -0.51 |
| 1a7c | TRP175 | -1.52 | -0.96 | -2.48 | 1.04 | -1.44 |
| 1a7c | VAL328 | -0.85 | -2.22 | -3.07 | 0.67 | -2.40 |
| 1a7c | ASN329 | -1.20 | -0.34 | -1.54 | 0.34 | -1.20 |