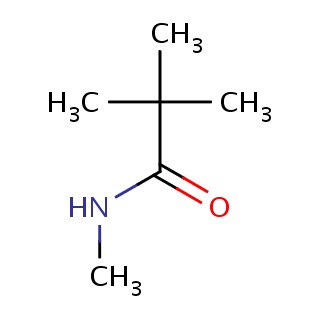
Binding information for 4r5v_ligand_2_0.mol2(FDBF00031)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4r5v_ligand_2_0.mol2 | 4r5v | 1 | -6.31 | CNC(=O)C(C)(C)C | 8 |
Structure and binding mode of 4r5v_ligand_2_0.mol2(FDBF00031)
Important binding residues for 4r5v_ligand_2_0.mol2(FDBF00031)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4r5v | VAL459 | -0.80 | -1.15 | -1.95 | 0.74 | -1.21 |
| 4r5v | GLY460 | -0.66 | -2.96 | -3.62 | 1.71 | -1.91 |
| 4r5v | ALA461 | -0.83 | -1.04 | -1.87 | 0.76 | -1.12 |
| 4r5v | VAL493 | -0.66 | -0.11 | -0.77 | 0.06 | -0.70 |
| 4r5v | HIS496 | -1.15 | 0.01 | -1.14 | 0.54 | -0.61 |
| 4r5v | GLU519 | -0.21 | -3.66 | -3.87 | 2.27 | -1.61 |
| 4r5v | TYR580 | -1.15 | 1.18 | 0.03 | -0.89 | -0.87 |