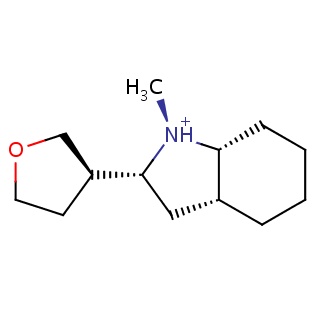
Binding information for 4gmc_ligand_1_1.mol2(FDBF01693)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4gmc_ligand_1_1.mol2 | 4gmc | 0.688525 | -6.85 | C[N@H+]1[C@@H]2CCCC[C@@H]2C[C@@H]1[C@H]1CCOC1 | 15 |
Structure and binding mode of 4gmc_ligand_1_1.mol2(FDBF01693)
Important binding residues for 4gmc_ligand_1_1.mol2(FDBF01693)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4gmc | LEU392 | -0.66 | -0.08 | -0.74 | 0.19 | -0.55 |
| 4gmc | ALA393 | -0.60 | -0.27 | -0.87 | 0.23 | -0.64 |
| 4gmc | ALA396 | -1.07 | -0.05 | -1.12 | -0.18 | -1.30 |
| 4gmc | LEU492 | -1.22 | -0.63 | -1.85 | 1.30 | -0.55 |
| 4gmc | VAL494 | -1.91 | -0.65 | -2.56 | 0.11 | -2.44 |
| 4gmc | PRO495 | -0.73 | 0.68 | -0.05 | -0.36 | -0.41 |
| 4gmc | TRP500 | -0.72 | 0.01 | -0.71 | 0.17 | -0.54 |