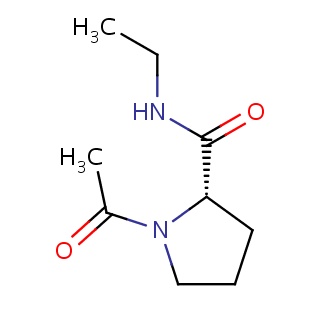
Binding information for 1ad8_ligand_4_148.mol2(FDBF00035)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1ad8_ligand_4_148.mol2 | 1ad8 | 1 | -6.62 | C1N(C(=O)C)[C@@H](CC1)C(=O)NCC | 13 |
Structure and binding mode of 1ad8_ligand_4_148.mol2(FDBF00035)
Important binding residues for 1ad8_ligand_4_148.mol2(FDBF00035)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1ad8 | HIS57 | -1.21 | 0.18 | -1.03 | 0.60 | -0.44 |
| 1ad8 | TYR60A | -1.04 | -0.40 | -1.44 | 0.85 | -0.59 |
| 1ad8 | TRP60D | -1.53 | -0.67 | -2.2 | 0.99 | -1.21 |
| 1ad8 | LEU99 | -0.76 | 0.13 | -0.63 | -0.16 | -0.80 |
| 1ad8 | SER214 | -0.97 | -2.61 | -3.58 | 2.32 | -1.26 |
| 1ad8 | TRP215 | -1.40 | -2.48 | -3.88 | 1.49 | -2.39 |
| 1ad8 | GLY216 | -0.82 | -0.13 | -0.95 | 0.13 | -0.82 |