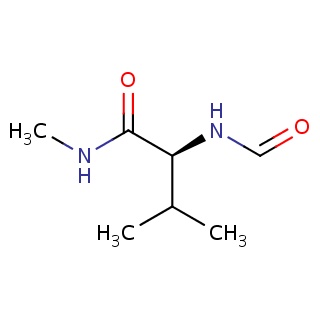
Binding information for 1ec1_ligand_3_885.mol2(FDBF00039)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1ec1_ligand_3_885.mol2 | 1ec1 | 1 | -6.61 | C(C)(C)[C@@H](C(=O)NC)NC=O | 11 |
Structure and binding mode of 1ec1_ligand_3_885.mol2(FDBF00039)
Important binding residues for 1ec1_ligand_3_885.mol2(FDBF00039)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1ec1 | GLY27 | -0.73 | -1.03 | -1.76 | 1.01 | -0.74 |
| 1ec1 | ALA28 | -1.51 | -1.35 | -2.86 | 0.62 | -2.24 |
| 1ec1 | ASP29 | -1.11 | -0.56 | -1.67 | -0.03 | -1.70 |
| 1ec1 | ILE47 | -0.83 | 0.56 | -0.27 | -0.38 | -0.66 |
| 1ec1 | GLY48 | -0.69 | -3.87 | -4.56 | 2.89 | -1.66 |
| 1ec1 | GLY49 | -0.76 | -2.01 | -2.77 | 1.44 | -1.34 |
| 1ec1 | ILE50 | -0.30 | -0.34 | -0.64 | 0.28 | -0.36 |
| 1ec1 | ILE84 | -0.62 | 0.01 | -0.61 | -0.02 | -0.63 |
| 1ec1 | ILE150 | -0.94 | -0.46 | -1.4 | 0.44 | -0.96 |