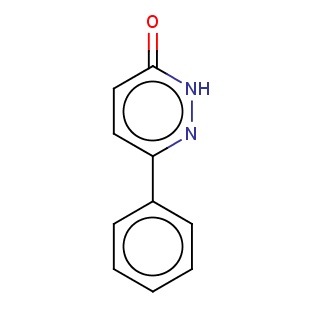
Binding information for 1xor_ligand_1_0.mol2(FDBF01829)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1xor_ligand_1_0.mol2 | 1xor | 0.494737 | -7.64 | O=c1[nH]nc(cc1)c1ccccc1 | 13 |
Structure and binding mode of 1xor_ligand_1_0.mol2(FDBF01829)
Important binding residues for 1xor_ligand_1_0.mol2(FDBF01829)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1xor | TYR159 | -2.06 | -0.67 | -2.73 | 1.79 | -0.94 |
| 1xor | ASP201 | -0.36 | 8.81 | 8.45 | -11.73 | -3.27 |
| 1xor | ASP318 | -0.86 | 2.61 | 1.75 | -5.25 | -3.51 |
| 1xor | LEU319 | -0.80 | 0.08 | -0.72 | -0.12 | -0.84 |
| 1xor | ILE336 | -1.75 | -0.00 | -1.75 | -0.34 | -2.10 |
| 1xor | PHE340 | -0.64 | 0.23 | -0.41 | 0.08 | -0.33 |
| 1xor | PHE372 | -1.58 | 0.52 | -1.06 | 0.11 | -0.95 |