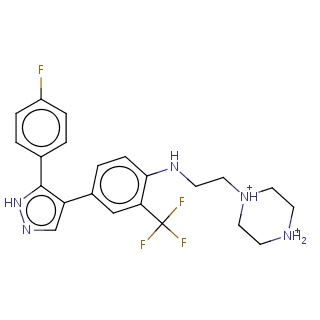
Binding information for 4xsz_ligand.mol2(FDBF01830)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4xsz_ligand.mol2 | 4xsz | 0.496296 | -9.14 | c1(F)ccc(cc1)c1c(c2ccc(NCC[NH+]3CC[NH2+]CC3)c(c2)C(F)(F)F)cn[nH]1 | 32 |
Structure and binding mode of 4xsz_ligand.mol2(FDBF01830)
Important binding residues for 4xsz_ligand.mol2(FDBF01830)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4xsz | ASP443 | -0.03 | -27.11 | -27.14 | 26.81 | -0.32 |
| 4xsz | ASP444 | 0.51 | -72.50 | -71.99 | 69.03 | -2.96 |
| 4xsz | GLU546 | -0.06 | -62.89 | -62.95 | 62.48 | -0.48 |
| 4xsz | HIS551 | -1.60 | 8.03 | 6.43 | -7.51 | -1.08 |
| 4xsz | PRO552 | -2.99 | 3.99 | 1 | -3.97 | -2.98 |
| 4xsz | GLU611 | -0.03 | -47.49 | -47.52 | 47.14 | -0.38 |
| 4xsz | ARG637 | -0.72 | 23.98 | 23.26 | -23.58 | -0.32 |
| 4xsz | SER642 | -1.90 | -3.46 | -5.36 | 3.03 | -2.32 |
| 4xsz | LYS749 | -1.90 | 52.01 | 50.11 | -50.64 | -0.53 |
| 4xsz | PRO750 | -1.34 | 3.18 | 1.84 | -3.24 | -1.40 |
| 4xsz | ASP751 | -0.28 | -35.66 | -35.94 | 35.54 | -0.40 |
| 4xsz | ILE755 | -1.34 | 0.78 | -0.56 | -0.93 | -1.48 |
| 4xsz | LEU770 | -0.94 | 0.03 | -0.91 | 0.37 | -0.53 |
| 4xsz | PHE773 | -1.58 | 0.21 | -1.37 | -0.04 | -1.41 |
| 4xsz | ILE774 | -1.49 | 0.08 | -1.41 | -0.08 | -1.50 |