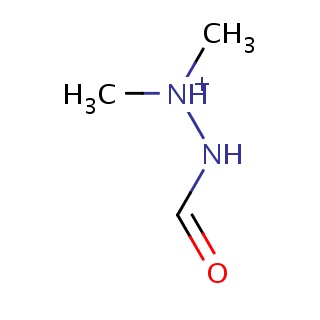
Binding information for 2cem_ligand_3_164.mol2(FDBF01907)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2cem_ligand_3_164.mol2 | 2cem | 0.363636 | -5.32 | C[NH+](NC=O)C | 6 |
Structure and binding mode of 2cem_ligand_3_164.mol2(FDBF01907)
Important binding residues for 2cem_ligand_3_164.mol2(FDBF01907)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2cem | GLY27 | -0.40 | -4.81 | -5.21 | 4.38 | -0.83 |
| 2cem | ALA28 | -0.54 | 1.15 | 0.61 | -1.02 | -0.41 |
| 2cem | ASP29 | -0.10 | -19.59 | -19.69 | 19.37 | -0.32 |
| 2cem | GLY49 | -0.43 | 1.12 | 0.69 | -1.26 | -0.57 |
| 2cem | ILE50 | -0.37 | 0.36 | -0.01 | -0.34 | -0.35 |
| 2cem | LEU123 | -0.25 | -19.12 | -19.37 | 18.98 | -0.40 |
| 2cem | ASP125 | -0.68 | -22.68 | -23.36 | 22.07 | -1.29 |
| 2cem | ILE150 | -0.55 | 1.04 | 0.49 | -1.20 | -0.70 |
| 2cem | ILE184 | -0.55 | 1.15 | 0.6 | -1.20 | -0.61 |