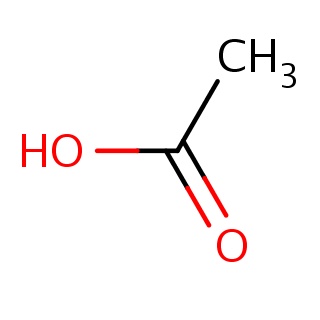
Binding information for 1r9l_ligand_frag_1.mol2(FDBF00004)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1r9l_ligand_frag_1.mol2 | 1r9l | 1 | -6.04 | CC(=O)O | 4 |
Structure and binding mode of 1r9l_ligand_frag_1.mol2(FDBF00004)
Important binding residues for 1r9l_ligand_frag_1.mol2(FDBF00004)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1r9l | PRO67 | -0.02 | -18.04 | -18.06 | 17.74 | -0.31 |
| 1r9l | LEU68 | -0.63 | -1.11 | -1.74 | 0.12 | -1.62 |
| 1r9l | HIS69 | 0.51 | -10.20 | -9.69 | 6.39 | -3.31 |
| 1r9l | CYS136 | -0.07 | -21.59 | -21.66 | 21.12 | -0.55 |
| 1r9l | GLY139 | -0.05 | -20.05 | -20.1 | 19.78 | -0.32 |
| 1r9l | TRP140 | -1.05 | -5.57 | -6.62 | 2.21 | -4.41 |
| 1r9l | GLY141 | 0.29 | -9.82 | -9.53 | 4.69 | -4.84 |
| 1r9l | CYS142 | 1.77 | -11.33 | -9.56 | 6.05 | -3.51 |