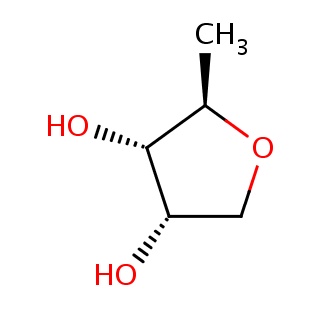
Binding information for 2xaf_ligand_1_3.mol2(FDBF00053)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2xaf_ligand_1_3.mol2 | 2xaf | 1 | -6.58 | [C@@H]1(OC[C@@H]([C@@H]1O)O)C | 8 |
Structure and binding mode of 2xaf_ligand_1_3.mol2(FDBF00053)
Important binding residues for 2xaf_ligand_1_3.mol2(FDBF00053)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2xaf | GLY285 | -0.67 | 0.64 | -0.03 | -1.01 | -1.04 |
| 2xaf | GLU308 | 2.99 | -20.29 | -17.3 | 14.31 | -2.99 |
| 2xaf | ALA309 | -0.43 | 0.54 | 0.11 | -0.98 | -0.87 |
| 2xaf | ARG310 | -0.60 | -4.74 | -5.34 | 1.72 | -3.63 |
| 2xaf | VAL313 | -0.12 | 0.11 | -0.01 | -0.46 | -0.47 |
| 2xaf | GLY314 | -0.22 | -0.25 | -0.47 | -0.08 | -0.56 |
| 2xaf | GLY315 | -0.54 | -0.24 | -0.78 | 0.38 | -0.40 |
| 2xaf | LEU625 | -0.50 | -0.17 | -0.67 | -0.01 | -0.68 |
| 2xaf | PRO626 | -0.68 | 0.07 | -0.61 | 0.10 | -0.51 |
| 2xaf | TRP756 | -0.95 | -0.39 | -1.34 | 0.71 | -0.63 |