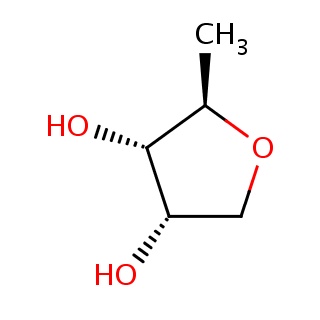
Binding information for 1h2t_ligand_1_10.mol2(FDBF00053)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1h2t_ligand_1_10.mol2 | 1h2t | 1 | -6.54 | [C@@H]1(OC[C@@H]([C@@H]1O)O)C | 8 |
Structure and binding mode of 1h2t_ligand_1_10.mol2(FDBF00053)
Important binding residues for 1h2t_ligand_1_10.mol2(FDBF00053)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1h2t | TYR43 | -0.80 | -0.43 | -1.23 | 0.52 | -0.71 |
| 1h2t | PHE83 | -1.24 | 0.27 | -0.97 | 0.13 | -0.84 |
| 1h2t | PHE85 | -0.77 | 0.68 | -0.09 | -0.36 | -0.45 |
| 1h2t | ARG123 | -0.70 | -13.41 | -14.11 | 5.63 | -8.47 |
| 1h2t | ARG127 | -1.73 | 3.05 | 1.32 | -2.11 | -0.78 |