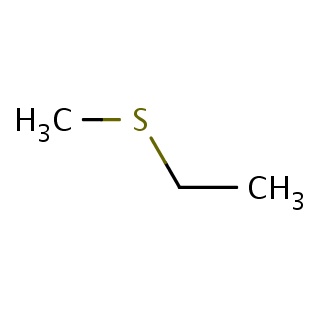
Binding information for 1pg2_ligand_2_2.mol2(FDBF00055)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1pg2_ligand_2_2.mol2 | 1pg2 | 1 | -5.86 | CCSC | 4 |
Structure and binding mode of 1pg2_ligand_2_2.mol2(FDBF00055)
Important binding residues for 1pg2_ligand_2_2.mol2(FDBF00055)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1pg2 | ALA12 | -0.75 | -0.84 | -1.59 | 0.25 | -1.33 |
| 1pg2 | LEU13 | -1.68 | -0.93 | -2.61 | 1.37 | -1.24 |
| 1pg2 | TRP253 | -1.04 | -0.24 | -1.28 | 0.81 | -0.48 |
| 1pg2 | ALA256 | -0.78 | 0.12 | -0.66 | -0.16 | -0.82 |
| 1pg2 | PRO257 | -0.68 | -0.05 | -0.73 | 0.20 | -0.53 |
| 1pg2 | ILE297 | -0.62 | 0.63 | 0.01 | -0.44 | -0.43 |
| 1pg2 | HIS301 | -0.65 | -0.84 | -1.49 | 1.03 | -0.46 |