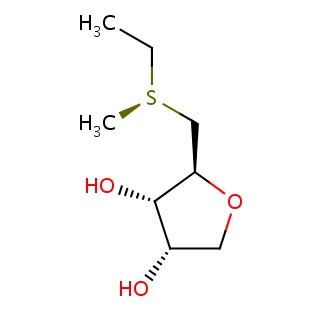
Binding information for 2pt9_ligand_4_10.mol2(FDBF00058)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2pt9_ligand_4_10.mol2 | 2pt9 | 1 | -6.91 | [C@@H]1(OC[C@@H]([C@@H]1O)O)C[S@@](C)CC | 12 |
Structure and binding mode of 2pt9_ligand_4_10.mol2(FDBF00058)
Important binding residues for 2pt9_ligand_4_10.mol2(FDBF00058)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2pt9 | GLN72 | -0.60 | -14.97 | -15.57 | 13.15 | -2.43 |
| 2pt9 | LEU86 | -0.39 | -19.05 | -19.44 | 18.81 | -0.62 |
| 2pt9 | LEU88 | -0.94 | -16.26 | -17.2 | 16.29 | -0.92 |
| 2pt9 | LEU94 | -0.44 | -0.39 | -0.83 | 0.24 | -0.58 |
| 2pt9 | GLY124 | -0.90 | 2.70 | 1.8 | -2.30 | -0.51 |
| 2pt9 | GLY125 | -1.11 | 2.80 | 1.69 | -2.72 | -1.02 |
| 2pt9 | GLY126 | -0.93 | 3.34 | 2.41 | -3.03 | -0.62 |
| 2pt9 | GLU147 | 2.92 | -51.07 | -48.15 | 42.50 | -5.65 |
| 2pt9 | ILE148 | -0.55 | 1.43 | 0.88 | -1.76 | -0.87 |
| 2pt9 | VAL152 | -0.36 | 0.59 | 0.23 | -1.17 | -0.94 |
| 2pt9 | SER197 | -1.00 | -0.26 | -1.26 | 0.78 | -0.48 |
| 2pt9 | SER198 | -1.39 | -0.59 | -1.98 | 0.67 | -1.31 |