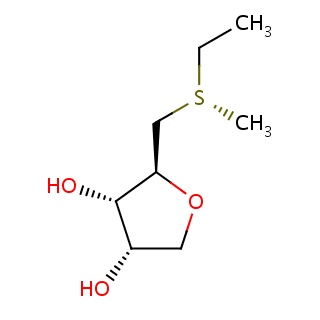
Binding information for 4ymg_ligand_4_10.mol2(FDBF00058)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 4ymg_ligand_4_10.mol2 | 4ymg | 1 | -6.64 | CC[S@](C)C[C@H]1OC[C@@H]([C@@H]1O)O | 12 |
Structure and binding mode of 4ymg_ligand_4_10.mol2(FDBF00058)
Important binding residues for 4ymg_ligand_4_10.mol2(FDBF00058)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 4ymg | GLY73 | -1.11 | -1.35 | -2.46 | 1.83 | -0.63 |
| 4ymg | CYS74 | -0.86 | 0.79 | -0.07 | -0.99 | -1.05 |
| 4ymg | TYR75 | -1.54 | 0.41 | -1.13 | -1.00 | -2.13 |
| 4ymg | GLU98 | 1.92 | -48.41 | -46.49 | 39.18 | -7.30 |
| 4ymg | TYR99 | -0.77 | 1.65 | 0.88 | -1.73 | -0.85 |
| 4ymg | SER100 | -0.46 | 0.97 | 0.51 | -0.91 | -0.40 |
| 4ymg | MET103 | -0.39 | 0.93 | 0.54 | -1.25 | -0.71 |
| 4ymg | ALA145 | -1.03 | -1.44 | -2.47 | 1.51 | -0.95 |