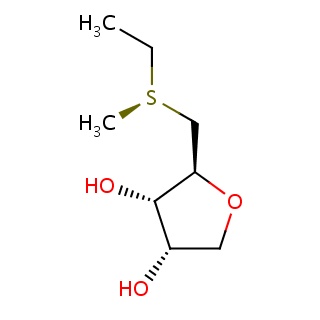
Binding information for 1nw5_ligand_4_10.mol2(FDBF00058)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 1nw5_ligand_4_10.mol2 | 1nw5 | 1 | -6.47 | [C@@H]1(OC[C@@H]([C@@H]1O)O)C[S@@](C)CC | 12 |
Structure and binding mode of 1nw5_ligand_4_10.mol2(FDBF00058)
Important binding residues for 1nw5_ligand_4_10.mol2(FDBF00058)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 1nw5 | PRO66 | -0.83 | -1.75 | -2.58 | 2.14 | -0.44 |
| 1nw5 | PRO67 | -1.25 | 0.12 | -1.13 | -0.47 | -1.60 |
| 1nw5 | HIS223 | -0.70 | 13.58 | 12.88 | -15.27 | -2.39 |
| 1nw5 | THR225 | -1.18 | -2.99 | -4.17 | 3.57 | -0.61 |
| 1nw5 | PHE250 | -1.78 | -3.88 | -5.66 | 3.82 | -1.83 |
| 1nw5 | ALA251 | -0.52 | -0.12 | -0.64 | 0.16 | -0.48 |
| 1nw5 | GLY252 | -0.85 | 2.18 | 1.33 | -2.12 | -0.79 |
| 1nw5 | ASP271 | 1.32 | -48.27 | -46.95 | 41.96 | -4.99 |
| 1nw5 | ALA272 | -0.41 | 1.82 | 1.41 | -1.92 | -0.51 |
| 1nw5 | ALA273 | -0.46 | 0.28 | -0.18 | -0.28 | -0.46 |