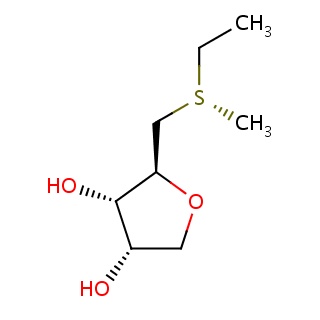
Binding information for 2zif_ligand_4_10.mol2(FDBF00058)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2zif_ligand_4_10.mol2 | 2zif | 1 | -6.30 | CC[S@](C)C[C@H]1OC[C@@H]([C@@H]1O)O | 12 |
Structure and binding mode of 2zif_ligand_4_10.mol2(FDBF00058)
Important binding residues for 2zif_ligand_4_10.mol2(FDBF00058)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2zif | SER47 | -0.54 | -3.73 | -4.27 | 3.95 | -0.32 |
| 2zif | TYR50 | -0.39 | 0.10 | -0.29 | -0.05 | -0.34 |
| 2zif | PHE243 | -0.78 | -4.21 | -4.99 | 4.14 | -0.85 |
| 2zif | ALA244 | -0.56 | -0.14 | -0.7 | 0.30 | -0.39 |
| 2zif | GLY245 | -0.98 | 1.60 | 0.62 | -1.79 | -1.17 |
| 2zif | GLU264 | -1.02 | -43.70 | -44.72 | 41.45 | -3.27 |
| 2zif | LEU265 | -0.63 | 2.14 | 1.51 | -2.32 | -0.81 |
| 2zif | VAL266 | -0.49 | 0.01 | -0.48 | -0.10 | -0.57 |
| 2zif | TYR269 | -0.69 | 15.43 | 14.74 | -15.09 | -0.35 |