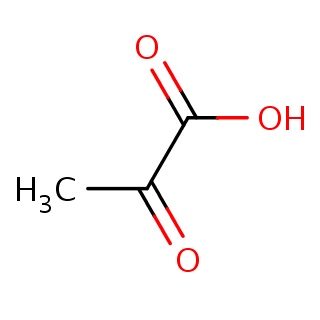
Binding information for 2hzl_ligand.mol2(FDBF00060)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2hzl_ligand.mol2 | 2hzl | 1 | -6.39 | C(=O)(O)C(=O)C | 7 |
Structure and binding mode of 2hzl_ligand.mol2(FDBF00060)
Important binding residues for 2hzl_ligand.mol2(FDBF00060)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2hzl | PHE40 | -0.33 | -0.98 | -1.31 | 0.99 | -0.32 |
| 2hzl | TYR100 | 1.11 | 5.16 | 6.27 | -7.95 | -1.68 |
| 2hzl | GLN156 | -0.01 | -20.62 | -20.63 | 16.99 | -3.64 |
| 2hzl | ARG177 | 2.01 | -72.55 | -70.54 | 57.77 | -12.77 |
| 2hzl | VAL178 | -0.10 | -1.98 | -2.08 | 1.48 | -0.59 |
| 2hzl | GLY179 | -0.41 | -3.12 | -3.53 | 2.62 | -0.92 |
| 2hzl | VAL216 | -0.68 | 0.34 | -0.34 | -0.37 | -0.71 |
| 2hzl | PRO243 | -0.12 | -18.72 | -18.84 | 18.45 | -0.39 |