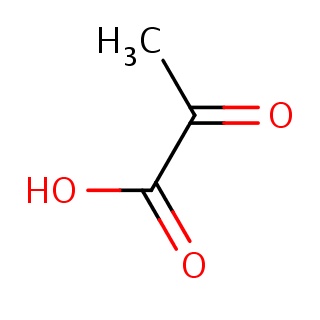
Binding information for 2xzw_ligand_1_0.mol2(FDBF00060)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2xzw_ligand_1_0.mol2 | 2xzw | 1 | -6.23 | CC(=O)C(=O)O | 6 |
Structure and binding mode of 2xzw_ligand_1_0.mol2(FDBF00060)
Important binding residues for 2xzw_ligand_1_0.mol2(FDBF00060)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2xzw | PHE36 | -0.70 | -5.75 | -6.45 | 4.06 | -2.38 |
| 2xzw | GLY37 | -0.40 | -6.49 | -6.89 | 3.83 | -3.06 |
| 2xzw | GLN39 | -1.76 | 4.27 | 2.51 | -5.29 | -2.78 |
| 2xzw | LYS40 | -0.64 | -21.60 | -22.24 | 20.33 | -1.90 |
| 2xzw | GLY41 | 1.21 | -6.85 | -5.64 | 4.60 | -1.04 |
| 2xzw | LEU56 | -0.45 | -0.44 | -0.89 | 0.42 | -0.46 |
| 2xzw | ILE86 | -0.96 | -2.88 | -3.84 | 2.55 | -1.29 |