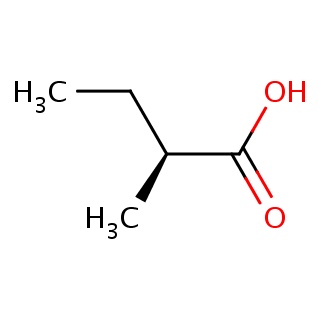
Binding information for 2i4j_ligand_1_0.mol2(FDBF00061)
| FRAGNAME | PDBID | SIMILIRITY | XSCORE | SMILE | HAC |
|---|---|---|---|---|---|
| 2i4j_ligand_1_0.mol2 | 2i4j | 1 | -6.37 | [C@H](C(=O)O)(C)CC | 7 |
Structure and binding mode of 2i4j_ligand_1_0.mol2(FDBF00061)
Important binding residues for 2i4j_ligand_1_0.mol2(FDBF00061)
| PDBID | RESIDUE | ΔEvdw | ΔEele | ΔEgas | ΔEsolv | ΔEgb |
|---|---|---|---|---|---|---|
| 2i4j | PHE282 | -0.61 | 1.24 | 0.63 | -0.99 | -0.37 |
| 2i4j | CYS285 | -0.42 | -0.36 | -0.78 | 0.47 | -0.31 |
| 2i4j | GLN286 | -0.88 | -0.80 | -1.68 | 1.19 | -0.48 |
| 2i4j | ARG288 | -0.05 | -14.43 | -14.48 | 14.13 | -0.36 |
| 2i4j | VAL322 | -0.03 | -17.14 | -17.17 | 16.87 | -0.31 |
| 2i4j | HIS323 | 1.86 | -13.13 | -11.27 | 6.81 | -4.46 |
| 2i4j | LYS367 | -0.04 | -35.75 | -35.79 | 35.37 | -0.42 |
| 2i4j | HIS449 | 1.16 | -27.98 | -26.82 | 24.27 | -2.56 |
| 2i4j | LEU469 | -0.39 | -0.50 | -0.89 | 0.24 | -0.65 |